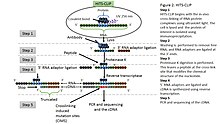
Protein – RNA interactions: Analysis of iCLIP-seq data FU Berlin Seminar RNA Bioinformatics, WS 14/15 Sabrina Krakau
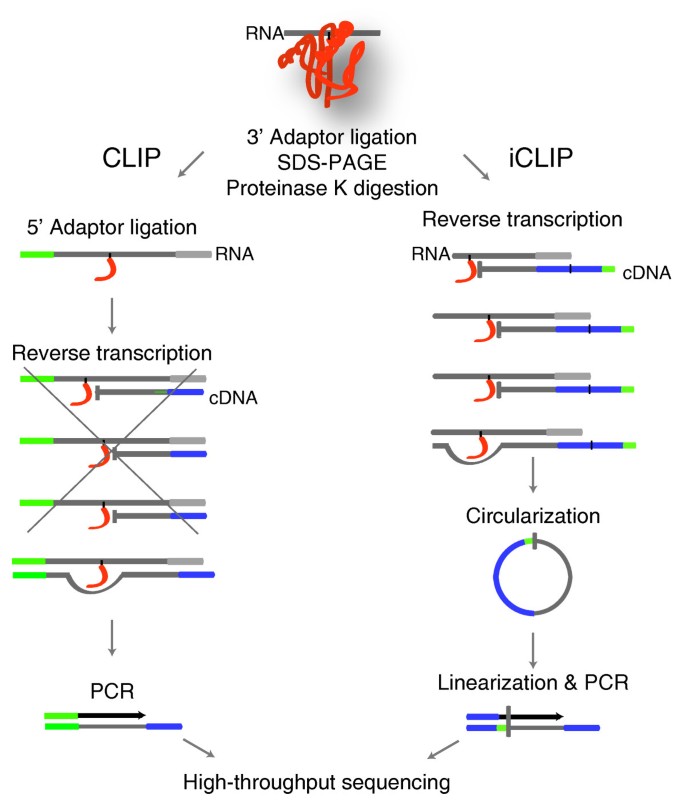
Analysis of CLIP and iCLIP methods for nucleotide-resolution studies of protein-RNA interactions | Genome Biology | Full Text
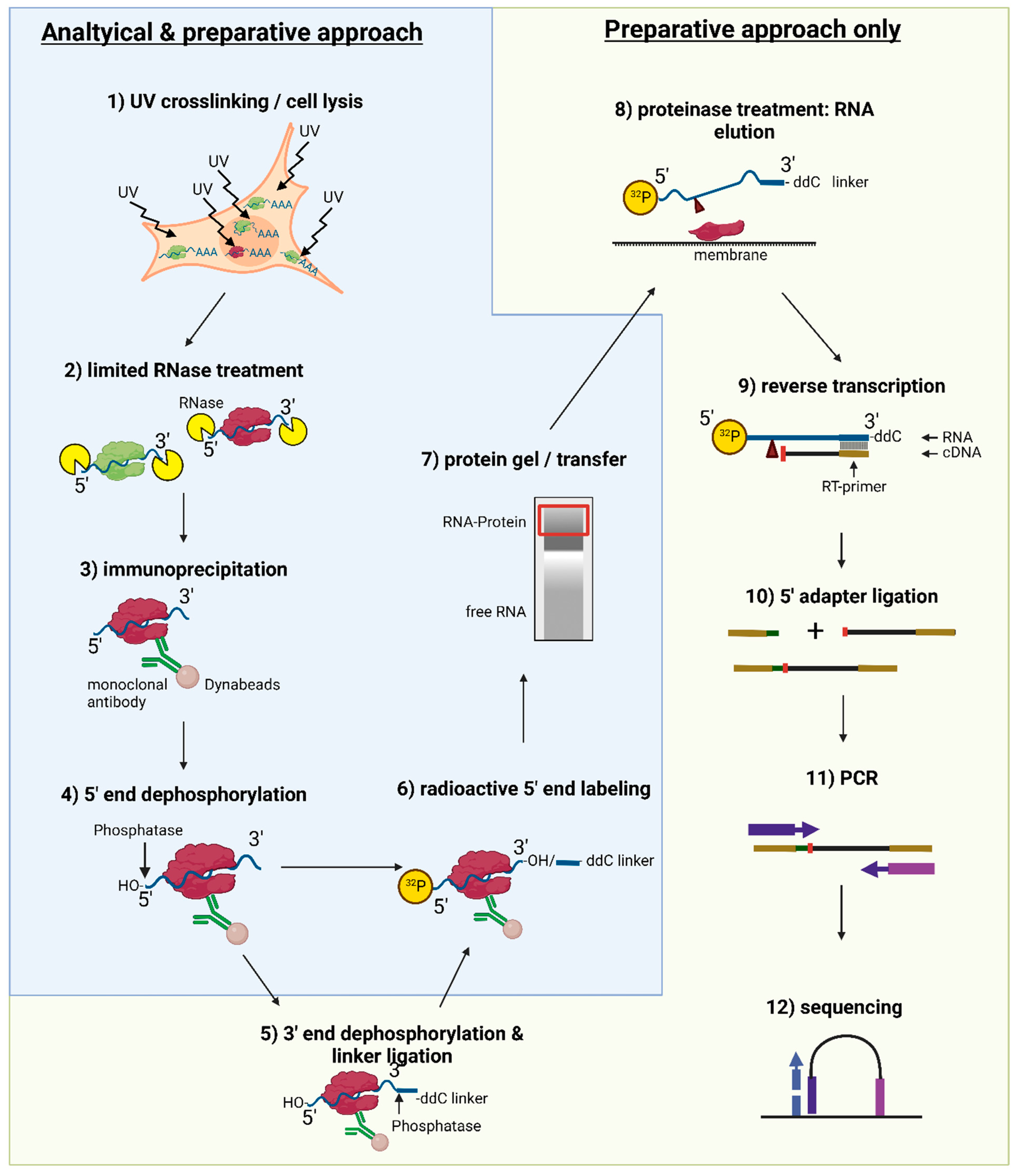
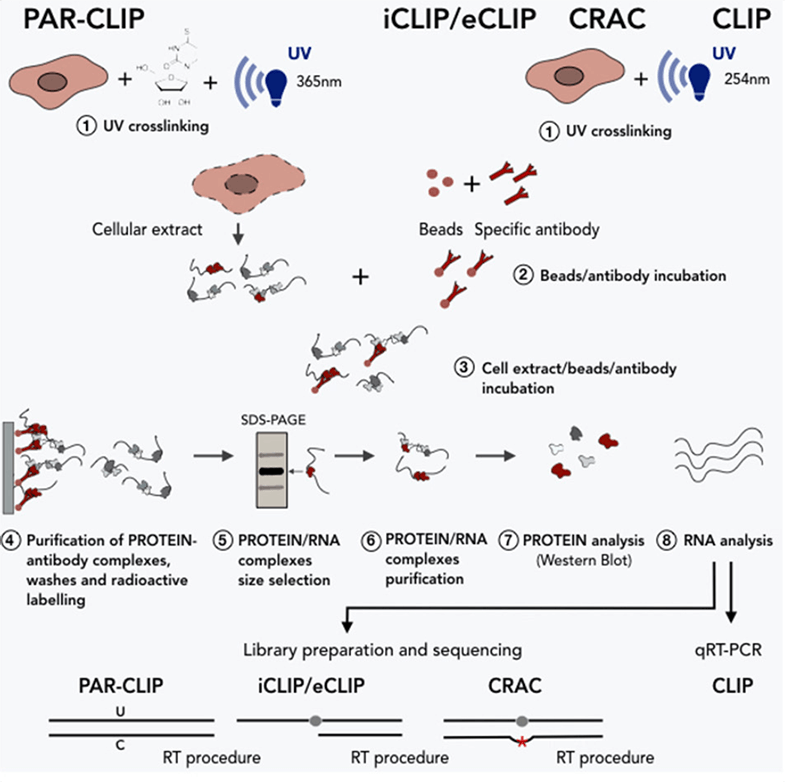
MPs | Free Full-Text | Studying miRNA–mRNA Interactions: An Optimized CLIP-Protocol for Endogenous Ago2-Protein
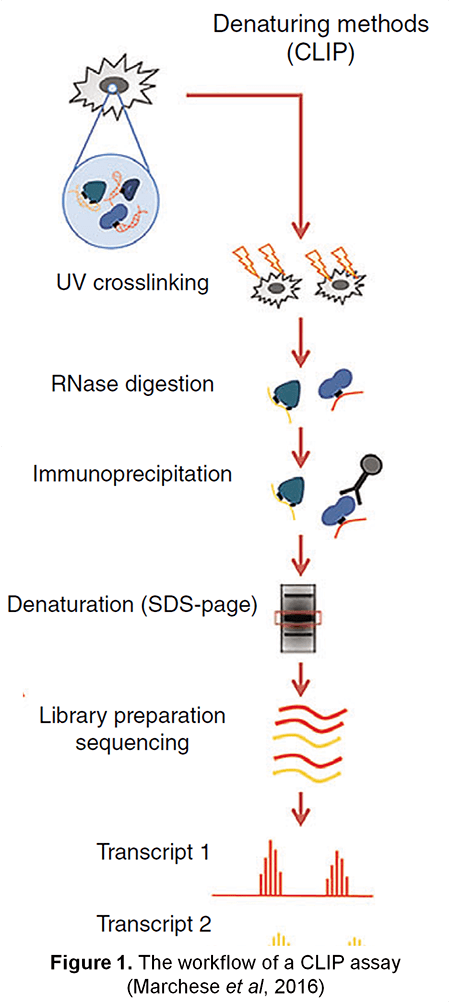
![PDF] CLIP: a method for identifying protein-RNA interaction sites in living cells. | Semantic Scholar PDF] CLIP: a method for identifying protein-RNA interaction sites in living cells. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/2f02e3c59d9a653227f0ccc71a5ccb088e0214ec/3-Figure1-1.png)






-2.jpg)
-3.jpg)


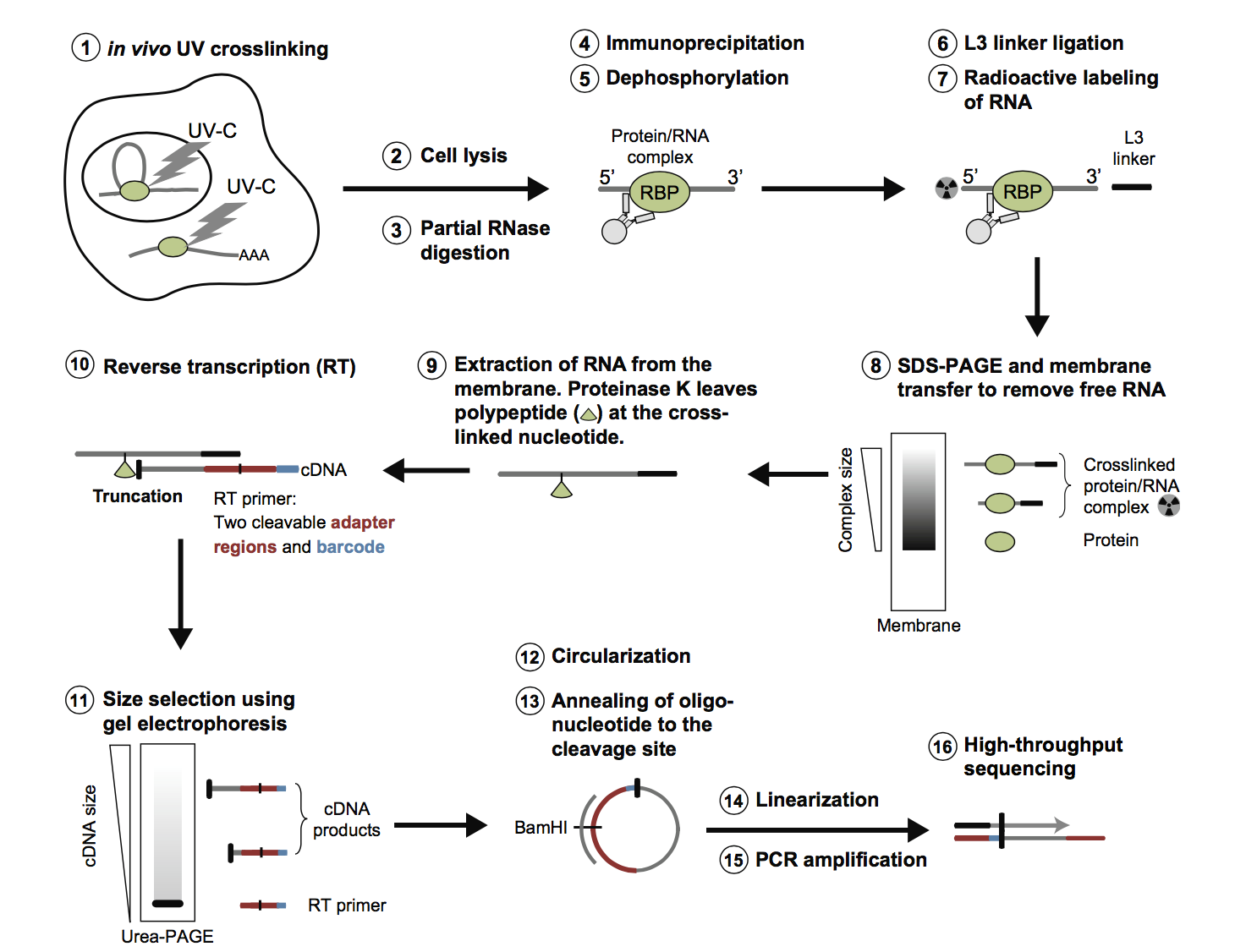
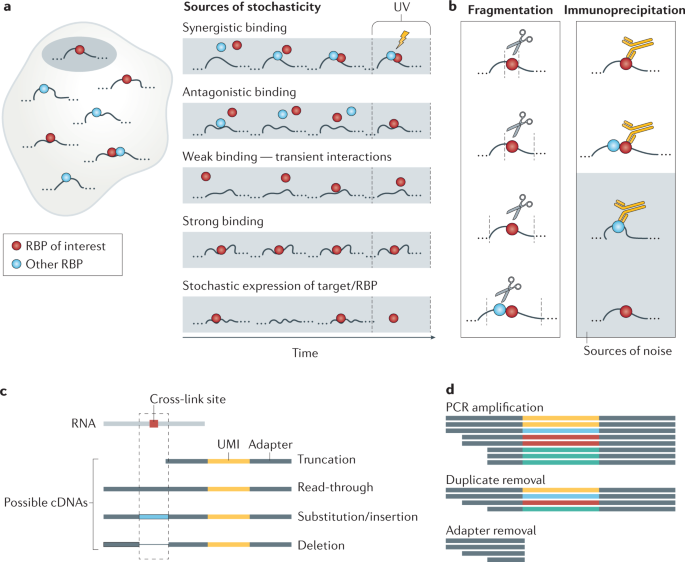
![PDF] The Future of Cross-Linking and Immunoprecipitation (CLIP). | Semantic Scholar PDF] The Future of Cross-Linking and Immunoprecipitation (CLIP). | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/f5ac369792eba3213c7d855f6e121dfcd9c4be6e/3-Figure1-1.png)